Products
BESPOKE SERVICE
Product Customization
Whether you need a bespoke antigen, a high-activity enzyme, or a tweak to a catalog reagent, we can help. Our team screens genes, fine-tunes fermentations from pilot to production scale, and purifies at up to 5,000L batches – delivering validated, assay-ready biomolecules with secure, repeatable supply.
Read more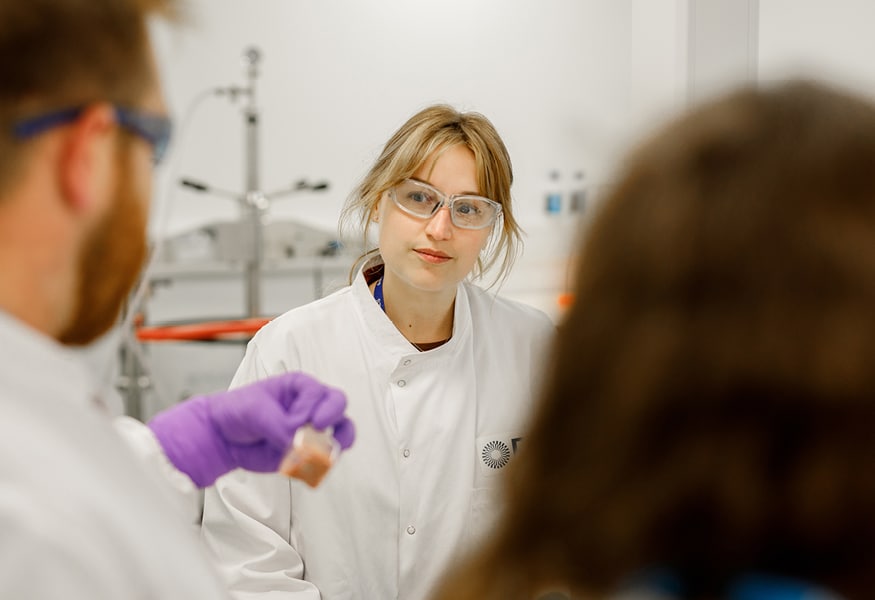
Sustainability
Advancing health, sustainably
We care passionately about helping you improve health and save lives. That passion extends to the planet. EcoVadis Silver already places us in the top 15% for sustainability, but we’re pushing further every day to become your most sustainable partner. Join us on the journey.
Our sustainability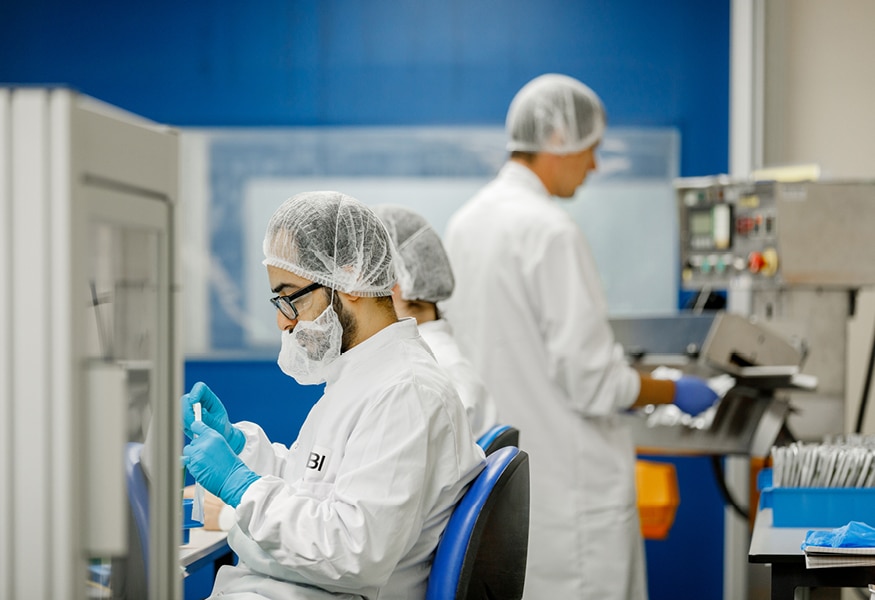
BBI AT A GLANCE
Certified
annual glucose strips powered by BBI enzymes
annual lateral flow test powered by BBI gold
Ecovadis rating
Recently Viewed
GET IN TOUCH
Still need help? Talk to our team
Our customer service team is on hand to help with any questions you have or support you need.
And if the issue turns technical, they’ll bring in our expert scientists to troubleshoot side-by-side with you and keep your project moving.
Contact us